19 results
Clostridioides difficile infection surveillance in intensive care units and oncology wards using machine learning
- Erkin Ötleş, Emily A. Balczewski, Micah Keidan, Jeeheh Oh, Alieysa Patel, Vincent B. Young, Krishna Rao, Jenna Wiens
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 44 / Issue 11 / November 2023
- Published online by Cambridge University Press:
- 24 April 2023, pp. 1776-1781
- Print publication:
- November 2023
-
- Article
-
- You have access Access
- Open access
- HTML
- Export citation
-
Objective:
Screening individuals admitted to the hospital for Clostridioides difficile presents opportunities to limit transmission and hospital-onset C. difficile infection (HO-CDI). However, detection from rectal swabs is resource intensive. In contrast, machine learning (ML) models may accurately assess patient risk without significant resource usage. In this study, we compared the effectiveness of swab surveillance to daily risk estimates produced by an ML model to identify patients who will likely develop HO-CDI in the intensive care unit (ICU) setting.
Design:A prospective cohort study was conducted with patient carriage of toxigenic C. difficile identified by rectal swabs analyzed by anaerobic culture and polymerase chain reaction (PCR). A previously validated ML model using electronic health record data generated daily risk of HO-CDI for every patient. Swab results and risk predictions were compared to the eventual HO-CDI status.
Patients:Adult inpatient admissions taking place in University of Michigan Hospitals’ medical and surgical intensive care units and oncology wards between June 6th and October 8th, 2020.
Results:In total, 2,979 admissions, representing 2,044 patients, were observed over the course of the study period, with 39 admissions developing HO-CDIs. Swab surveillance identified 9 true-positive and 87 false-positive HO-CDIs. The ML model identified 9 true-positive and 226 false-positive HO-CDIs; 8 of the true-positives identified by the model differed from those identified by the swab surveillance.
Conclusion:With limited resources, an ML model identified the same number of HO-CDI admissions as swab-based surveillance, though it generated more false-positives. The patients identified by the ML model were not yet colonized with C. difficile. Additionally, the ML model identifies at-risk admissions before disease onset, providing opportunities for prevention.
Prospective evaluation of data-driven models to predict daily risk of Clostridioides difficile infection at 2 large academic health centers – ERRATUM
- Meghana Kamineni, Erkin Ötleş, Jeeheh Oh, Krishna Rao, Vincent B. Young, Benjamin Y. Li, Lauren R. West, David C. Hooper, Erica S. Shenoy, John G. Guttag, Jenna Wiens, Maggie Makar
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 44 / Issue 10 / October 2023
- Published online by Cambridge University Press:
- 19 December 2022, p. 1705
- Print publication:
- October 2023
-
- Article
-
- You have access Access
- HTML
- Export citation
Prospective evaluation of data-driven models to predict daily risk of Clostridioides difficile infection at 2 large academic health centers
- Meghana Kamineni, Erkin Ötleş, Jeeheh Oh, Krishna Rao, Vincent B. Young, Benjamin Y. Li, Lauren R. West, David C. Hooper, Erica S. Shenoy, John G. Guttag, Jenna Wiens, Maggie Makar
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 44 / Issue 7 / July 2023
- Published online by Cambridge University Press:
- 19 September 2022, pp. 1163-1166
- Print publication:
- July 2023
-
- Article
- Export citation
-
Many data-driven patient risk stratification models have not been evaluated prospectively. We performed and compared the prospective and retrospective evaluations of 2 Clostridioides difficile infection (CDI) risk-prediction models at 2 large academic health centers, and we discuss the models’ robustness to data-set shifts.
The prescriber’s guide to classic MAO inhibitors (phenelzine, tranylcypromine, isocarboxazid) for treatment-resistant depression
- Vincent Van den Eynde, Wegdan R. Abdelmoemin, Magid M. Abraham, Jay D. Amsterdam, Ian M. Anderson, Chittaranjan Andrade, Glen B. Baker, Aartjan T.F. Beekman, Michael Berk, Tom K. Birkenhäger, Barry B. Blackwell, Pierre Blier, Marc B.J. Blom, Alexander J. Bodkin, Carlo I. Cattaneo, Bezalel Dantz, Jonathan Davidson, Boadie W. Dunlop, Ryan F. Estévez, Shalom S. Feinberg, John P.M. Finberg, Laura J. Fochtmann, David Gotlib, Andrew Holt, Thomas R. Insel, Jens K. Larsen, Rajnish Mago, David B. Menkes, Jonathan M. Meyer, David J. Nutt, Gordon Parker, Mark D. Rego, Elliott Richelson, Henricus G. Ruhé, Jerónimo Sáiz-Ruiz, Stephen M. Stahl, Thomas Steele, Michael E. Thase, Sven Ulrich, Anton J.L.M. van Balkom, Eduard Vieta, Ian Whyte, Allan H. Young, Peter K. Gillman
-
- Journal:
- CNS Spectrums / Volume 28 / Issue 4 / August 2023
- Published online by Cambridge University Press:
- 15 July 2022, pp. 427-440
-
- Article
-
- You have access Access
- Open access
- HTML
- Export citation
-
This article is a clinical guide which discusses the “state-of-the-art” usage of the classic monoamine oxidase inhibitor (MAOI) antidepressants (phenelzine, tranylcypromine, and isocarboxazid) in modern psychiatric practice. The guide is for all clinicians, including those who may not be experienced MAOI prescribers. It discusses indications, drug-drug interactions, side-effect management, and the safety of various augmentation strategies. There is a clear and broad consensus (more than 70 international expert endorsers), based on 6 decades of experience, for the recommendations herein exposited. They are based on empirical evidence and expert opinion—this guide is presented as a new specialist-consensus standard. The guide provides practical clinical advice, and is the basis for the rational use of these drugs, particularly because it improves and updates knowledge, and corrects the various misconceptions that have hitherto been prominent in the literature, partly due to insufficient knowledge of pharmacology. The guide suggests that MAOIs should always be considered in cases of treatment-resistant depression (including those melancholic in nature), and prior to electroconvulsive therapy—while taking into account of patient preference. In selected cases, they may be considered earlier in the treatment algorithm than has previously been customary, and should not be regarded as drugs of last resort; they may prove decisively effective when many other treatments have failed. The guide clarifies key points on the concomitant use of incorrectly proscribed drugs such as methylphenidate and some tricyclic antidepressants. It also illustrates the straightforward “bridging” methods that may be used to transition simply and safely from other antidepressants to MAOIs.
Self-blame in major depression: a randomised pilot trial comparing fMRI neurofeedback with self-guided psychological strategies
- Tanja Jaeckle, Steven C. R. Williams, Gareth J. Barker, Rodrigo Basilio, Ewan Carr, Kimberley Goldsmith, Alessandro Colasanti, Vincent Giampietro, Anthony Cleare, Allan H. Young, Jorge Moll, Roland Zahn
-
- Journal:
- Psychological Medicine / Volume 53 / Issue 7 / May 2023
- Published online by Cambridge University Press:
- 02 December 2021, pp. 2831-2841
-
- Article
-
- You have access Access
- Open access
- HTML
- Export citation
-
Background
Overgeneralised self-blame and worthlessness are key symptoms of major depressive disorder (MDD) and have previously been associated with self-blame-selective changes in connectivity between right superior anterior temporal lobe (rSATL) and subgenual frontal cortices. Another study showed that remitted MDD patients were able to modulate this neural signature using functional magnetic resonance imaging (fMRI) neurofeedback training, thereby increasing their self-esteem. The feasibility and potential of using this approach in symptomatic MDD were unknown.
MethodThis single-blind pre-registered randomised controlled pilot trial probed a novel self-guided psychological intervention with and without additional rSATL-posterior subgenual cortex (BA25) fMRI neurofeedback, targeting self-blaming emotions in people with insufficiently recovered MDD and early treatment-resistance (n = 43, n = 35 completers). Participants completed three weekly self-guided sessions to rebalance self-blaming biases.
ResultsAs predicted, neurofeedback led to a training-induced reduction in rSATL-BA25 connectivity for self-blame v. other-blame. Both interventions were safe and resulted in a 46% reduction on the Beck Depression Inventory-II, our primary outcome, with no group differences. Secondary analyses, however, revealed that patients without DSM-5-defined anxious distress showed a superior response to neurofeedback compared with the psychological intervention, and the opposite pattern in anxious MDD. As predicted, symptom remission was associated with increases in self-esteem and this correlated with the frequency with which participants employed the psychological strategies in daily life.
ConclusionsThese findings suggest that self-blame-rebalance neurofeedback may be superior over a solely psychological intervention in non-anxious MDD, although further confirmatory studies are needed. Simple self-guided strategies tackling self-blame were beneficial, but need to be compared against treatment-as-usual in further trials. https://doi.org/10.1186/ISRCTN10526888
Admission and Discharge Sampling Underestimates Multidrug-Resistant Organism (MDRO) Acquisition in an Intensive Care Unit
- Sarah Sansom, Michael Lin, Christine Fukuda, Teppei Shimasaki, Thelma Dangana, Nicholas Moore, Rachel Yelin, Yoona Rhee, Lina Tabith, Jianrong Sheng, Enrique Cornejo Cisneros, John Murray, Kyle Chang, Karen Lolans, Michelle Ariston, William Rotunno, Hazel Ramos, Haiying Li, Khaled Aboushaala, Naomi Iwai, Christine Bassis, Vincent Young, Mary Hayden
-
- Journal:
- Antimicrobial Stewardship & Healthcare Epidemiology / Volume 1 / Issue S1 / July 2021
- Published online by Cambridge University Press:
- 29 July 2021, p. s28
-
- Article
-
- You have access Access
- Open access
- Export citation
-
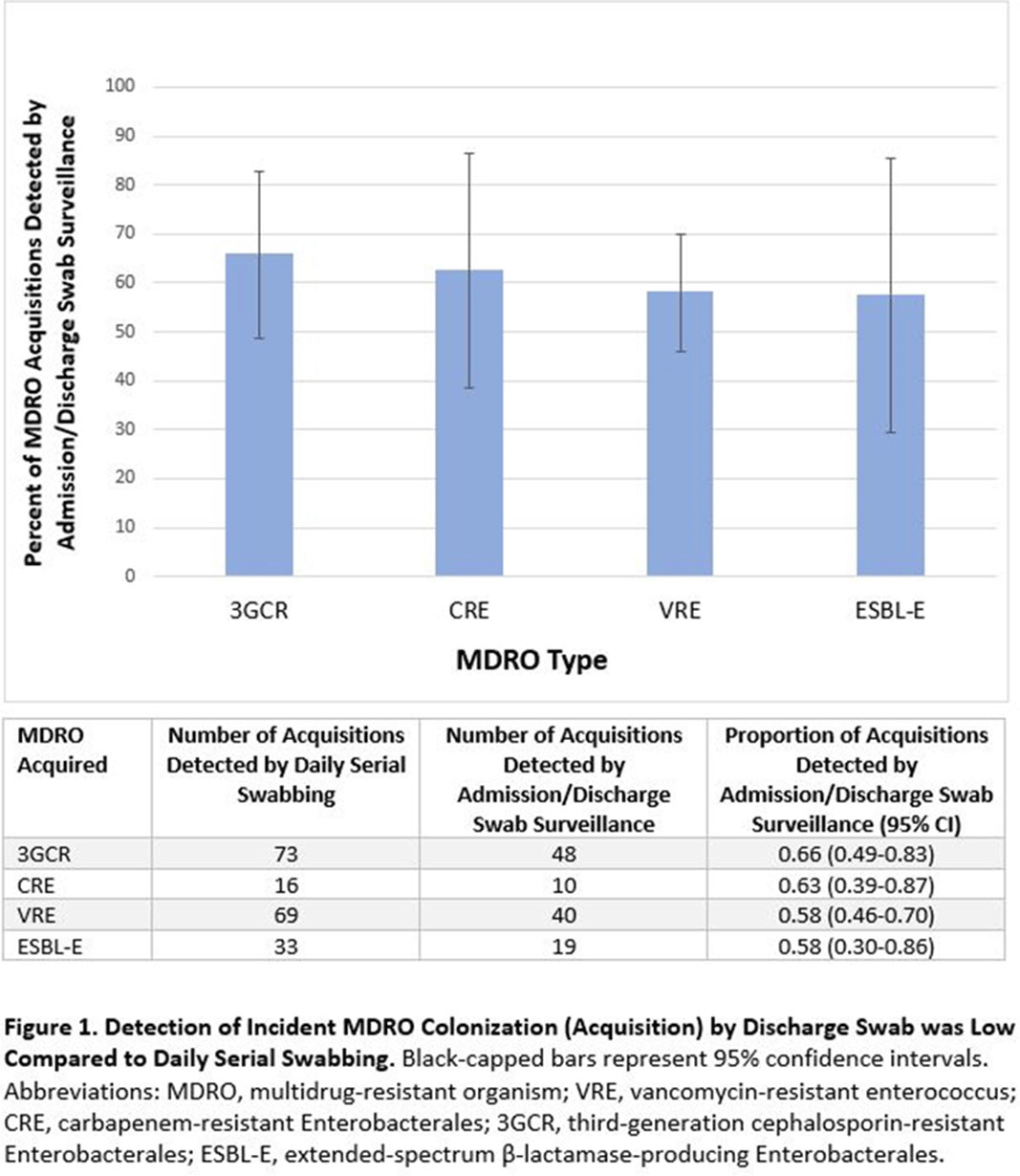
Background: Identification of hospitalized patients with enteric multidrug-resistant organism (MDRO) carriage, combined with implementation of targeted infection control interventions, may help reduce MDRO transmission. However, the optimal surveillance approach has not been defined. We sought to determine whether daily serial rectal surveillance for MDROs detects more incident cases (acquisition) of MDRO colonization in medical intensive care unit (MICU) patients than admission and discharge surveillance alone. Methods: Prospective longitudinal observational single-center study from January 11, 2017, to January 11, 2018. Inclusion criteria were ≥3 consecutive MICU days and ≥2 rectal or stool swabs per MICU admission. Daily rectal or stool swabs were collected from patients and cultured for MDROs, including vancomycin-resistant Enterococcus (VRE), carbapenem-resistant Enterobacterales (CRE), third-generation cephalosporin-resistant Enterobacterales (3GCR), and extended-spectrum β-lactamase–producing Enterobacterales (ESBL-E) (as a subset of 3GCR). MDRO detection at any time during the MICU stay was used to calculate prevalent colonization. Incident colonization (acquisition) was defined as new detection of an MDRO after at least 1 prior negative swab. We then determined the proportion of prevalent and incident cases detected by daily testing that were also detected when only first swabs (admission) and last swabs (discharge) were tested. Data were analyzed using SAS version 9.4 software. Results: In total, 939 MICU stays of 842 patients were analyzed. Patient characteristics were median age 64 years (interquartile range [IQR], 51–74), median MICU length of stay 5 days (IQR, 3–8), median number of samples per admission 3 (IQR, 2–5), and median Charlson index 4 (IQR, 2–7). Prevalent colonization with any MDRO was detected by daily swabbing in 401 stays (42.7%). Compared to daily serial swabbing, an admission- and discharge-only approach detected ≥86% of MDRO cases (ie, overall prevalent MDRO colonization). Detection of incident MDRO colonization by an admission- or discharge-only approach would have detected fewer cases than daily swabbing (Figure 1); ≥34% of total MDRO acquisitions would have been missed. Conclusions: Testing patients upon admission and discharge to an MICU may fail to detect MDRO acquisition in more than one-third of patients, thereby reducing the effectiveness of MDRO control programs that are targeted against known MDRO carriers. The poor performance of a single discharge swab may be due to intermittent or low-level MDRO shedding, inadequate sampling, or transient MDRO colonization. Additional research is needed to determine the optimal surveillance approach of enteric MDRO carriage.
Funding: No
Disclosures: None
Figure 1.
95332 Intestinal inflammation and altered gut microbiota associated with inflammatory bowel disease render mice susceptible to Clostridioides difficile colonization and infection
- Lisa Abernathy-Closedeline R. Barron, James M. George, Kimberly C. Vendrov, Peter D.R. Higgins, Ingrid L. Bergin, Vincent B. Young
-
- Journal:
- Journal of Clinical and Translational Science / Volume 5 / Issue s1 / March 2021
- Published online by Cambridge University Press:
- 31 March 2021, p. 90
-
- Article
-
- You have access Access
- Open access
- Export citation
-
ABSTRACT IMPACT: Use of this novel murine model of inflammatory bowel disease (IBD) and C. difficile infection (CDI) will aid in developing new clinical approaches to predict, diagnose, and treat CDI in the IBD population. OBJECTIVES/GOALS: IBD is associated with intestinal inflammation and alterations of the gut microbiota, both of which can diminish colonization resistance to C. difficile. Here, we sought to determine if IBD is sufficient to render mice susceptible to C. difficile colonization and infection in the absence of other perturbations, such as antibiotic treatment. METHODS/STUDY POPULATION: C57BL/6 IL-10-/- mice were colonized with Helicobacter hepaticus to trigger colonic inflammation akin to human IBD. Control mice, not colonized with H. hepaticus, were pretreated with the antibiotic cefoperazone to render the gut microbiota susceptible to CDI. Mice were then gavaged with spores of the toxigenic C. difficile strain VPI 10463 and monitored for C. difficile colonization and disease. The fecal microbiota at the time of C. difficile exposure was profiled by 16S rRNA gene sequencing and analyzed using mothur. Statistical analyses were performed using R. RESULTS/ANTICIPATED RESULTS: Mice with IBD harbored significantly distinct intestinal microbial communities compared to non-IBD controls at the time of C. difficile spore exposure (14 days post-IBD trigger). Mice with IBD were susceptible to persistent C. difficile colonization, while genetically identical non-IBD controls were resistant to C. difficile colonization. Concomitant IBD and CDI was associated with significantly worse clinical and intestinal disease than unaccompanied IBD. DISCUSSION/SIGNIFICANCE OF FINDINGS: Patients with IBD who develop concurrent CDI experience increased morbidity and mortality. These studies in a novel mouse model of IBD and CDI emphasize the dual importance of host responses and alterations of the gut microbiota in susceptibility to C. difficile colonization and infection in the setting of IBD.
54101 Characterizing Microbiota Features of Clostridioides difficile Infections
- Sarah Tomkovich, Krishna Rao, Vincent B. Young, Patrick D. Schloss
-
- Journal:
- Journal of Clinical and Translational Science / Volume 5 / Issue s1 / March 2021
- Published online by Cambridge University Press:
- 31 March 2021, p. 89
-
- Article
-
- You have access Access
- Open access
- Export citation
-
ABSTRACT IMPACT: Our goal is to identify bacterial biomarkers of adverse Clostridioides difficile infection outcomes OBJECTIVES/GOALS: We characterized microbiota features of Clostridioides difficile infections (CDIs) and will investigate the association between bacterial taxa and adverse outcomes, which includes severe and recurrent CDIs. METHODS/STUDY POPULATION: 1,517 stool samples were collected from patients diagnosed with a CDI at the University of Michigan along with 1,516 unformed and 910 formed stool control samples. We characterized the microbiota of the 3,943 stool samples by sequencing the V4 region of the 16S rRNA gene and used the Dirichlet Multinomial Mixtures method to cluster samples into community types. Severe CDI cases were defined using the Infectious Diseases Society of America criteria and recurrent CDIs were defined as CDIs that occurred within 2-12 weeks of the primary CDI. We will use machine learning to examine whether specific bacterial taxa can predict severe or recurrent CDIs. We will test 5 machine learning models with 80% training and 20% testing data split. RESULTS/ANTICIPATED RESULTS: Similar to findings from a previous study with 338 samples, we found there was no difference in diversity between CDI cases and unformed controls (Inverse Simpson index, p > 0.5) and samples from the 3 groups (CDIs, unformed controls, and formed controls) clustered into 12 community types. To investigate the bacterial taxa that are important for predicting adverse CDI outcomes, we will select the best machine learning model based on performance and training time and examine how much each feature contributes to performance. We anticipate the large number of CDI cases in our cohort and robust machine learning approaches will enable us to identify more bacteria associated with adverse outcomes compared to other studies that have attempted to predict CDI recurrence with fewer CDI cases. DISCUSSION/SIGNIFICANCE OF FINDINGS: Adverse CDI outcomes are a significant source of the morbidities, mortalities, and healthcare costs associated with CDIs. Identifying bacterial biomarkers of severe and recurrent CDIs could enhance our ability to stratify patients into risk groups and may lead to the development of more targeted therapeutics.
Genomic Epidemiology of Clostridioides difficile Sequence Types 1 and 2 Across Three US Medical Centers
- Arianna Miles-Jay, Vincent Young, Eric Pamer, Tor Savidge, Mini Kamboj, Kevin Garey, Evan Snitkin
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, p. s238
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background:Clostridioides difficile is a toxin-producing bacterium that is the foremost cause of healthcare-associated diarrhea in the United States. Recent epidemiologic and genomic evidence indicates that divergent C. difficile strains have varying propensities for transmission within healthcare settings. We investigated whether and how these differences are reflected in the genomic epidemiology of 2 common C. difficile strains—sequence type (ST) 1 (analogous to Ribotype 027) and ST2 (associated with Ribotypes 014/020)—across 3 geographically distinct US medical centers. Methods: Between 2011 and 2017, a convenience sample of ST1 and ST2 C. difficile clinical isolates were collected from 3 US sites: The University of Michigan Medical Center, Texas Medical Center Hospitals, and Memorial Sloan Kettering Cancer Center. Isolates underwent whole-genome sequencing and in silico multilocus sequence typing to verify strain types. Sequences were mapped to ST1 and ST2 reference genomes and single nucleotide variants (SNVs) were identified, filtered, and used to construct pairwise SNV distance matrices. A range of pairwise SNV distance thresholds were applied to assess genetic linkages consistent with recent transmission within ST1 compared to within ST2. Proportions of genetically linked isolates were compared using 2 tests. Results: We identified 200 ST1 and 188 ST2 isolates across the 3 collection sites. Overall, ST2 was more genetically diverse than ST1 (pairwise SNV distance range, 0–156 SNVs and 0–78 SNVs, respectively). ST2 isolates displayed significantly less evidence of recent transmission: 10 ST2 isolates (5.3%) were within 2 SNVs of another isolate compared to 88 (44%) ST1 isolates (P .001) (Fig. 1). As the SNV threshold increased to 5 and 10 SNVs, this trend was maintained (all P < .001). ST2 isolates were also more likely to be genetically linked to an isolate from a different collection site than ST1 isolates. Among isolates with genetic links to at least 1 other isolate at the 5 SNV and 10 SNV thresholds, 21 of 37 and 74 of 89 ST2 isolates (57%, 83%) were linked to an isolate from a different collection site, compared to 2 of 88 and 48 of 157 ST1 isolates (2% and 31%, respectively; both P < .001). Conclusions: Compared to C. difficile ST1 isolates, ST2 isolates displayed less evidence of recent healthcare transmission and were more likely to be genetically linked to isolates from divergent collection sites. Interpreting genetic linkages among C. difficile isolates requires an understanding of regional and strain-specific genetic diversity to avoid misattribution of genetic linkages to recent transmission.
Funding: None
Disclosures: None
Recurrent Clostridioides difficile infection can be predicted using inflammatory mediator and toxin activity levels
- Jonathan Motyka, Aline Penkevich, Vincent Young, Krishna Rao
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s77-s78
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background:Clostridioides difficile infection (CDI) frequently recurs after initial treatment. Predicting recurrent CDI (rCDI) early in the disease course can assist clinicians in their decision making and improve outcomes. However, predictions based on clinical criteria alone are not accurate and/or do not validate other results. Here, we tested the hypothesis that circulating and stool-derived inflammatory mediators predict rCDI. Methods: Consecutive subjects with available specimens at diagnosis were included if they tested positive for toxigenic C. difficile (+enzyme immunoassay [EIA] for glutamate dehydrogenase and toxins A/B, with reflex to PCR for the tcdB gene for discordants). Stool was thawed on ice, diluted 1:1 in PBS with protease inhibitor, centrifuged, and used immediately. A 17-plex panel of inflammatory mediators was run on a Luminex 200 machine using a custom antibody-linked bead array. Prior to analysis, all measurements were normalized and log-transformed. Stool toxin activity levels were quantified using a custom cell-culture assay. Recurrence was defined as a second episode of CDI within 100 days. Ordination characterized variation in the panel between outcomes, tested with a permutational, multivariate ANOVA. Machine learning via elastic net regression with 100 iterations of 5-fold cross validation selected the optimal model and the area under the receiver operator characteristic curve (AuROC) was computed. Sensitivity analyses excluding those that died and/or lived >100 km away were performed. Results: We included 186 subjects, with 95 women (51.1%) and average age of 55.9 years (±20). More patients were diagnosed by PCR than toxin EIA (170 vs 55, respectively). Death, rCDI, and no rCDI occurred in 32 (17.2%), 36 (19.4%), and 118 (63.4%) subjects, respectively. Ordination revealed that the serum panel was associated with rCDI (P = .007) but the stool panel was not. Serum procalcitonin, IL-8, IL-6, CCL5, and EGF were associated with recurrence. The machine-learning models using the serum panel predicted rCDI with AuROCs between 0.74 and 0.8 (Fig. 1). No stool inflammatory mediators independently predicted rCDI. However, stool IL-8 interacted with toxin activity to predict rCDI (Fig. 2). These results did not change significantly upon sensitivity analysis. Conclusions: A panel of serum inflammatory mediators predicted rCDI with up to 80% accuracy, but the stool panel alone was less successful. Incorporating toxin activity levels alongside inflammatory mediator measurements is a novel, promising approach to studying stool-derived biomarkers of rCDI. This approach revealed that stool IL-8 is a potential biomarker for rCDI. These results need to be confirmed both with a larger dataset and after adjustment for clinical covariates.
Funding: None
Disclosure: Vincent Young is a consultant for Bio-K+ International, Pantheryx, and Vedanta Biosciences.
Blind Spots in Methods Based on Cultivation and Metagenomic Sequencing for Surface Microbiomes in a Medical Intensive Care Unit
- Jiaxian Shen, Alexander McFarland, Ryan Blaustein, Mary Hayden, Vincent Young, Erica Hartmann
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 41 / Issue S1 / October 2020
- Published online by Cambridge University Press:
- 02 November 2020, pp. s141-s142
- Print publication:
- October 2020
-
- Article
-
- You have access Access
- Export citation
-
Background: Cultivation of targeted pathogens has been long recognized as a gold standard for healthcare surveillance. However, there is an emergent need to characterize all viable microorganisms in healthcare facilities to understand the role that both clinical and nonclinical microorganisms play in healthcare-associated infections. Metagenomic sequencing allows detection of entire microbial communities, in contrast to targeted identification by cultivation. Widespread application of metagenomic sequencing has been impeded in part because the sensitivity and specificity are unknown, which inhibits our ability to interpret results for risk assessment. To assess the impact of sample preparation methods on sensitivity and specificity, we compared several pretreatment steps followed by metagenomic sequencing, and we performed culture-based analyses. Methods: We collected 120 surface swabs from the medical intensive care unit at Rush University Medical Center, which we aggregated to create a representative microbiome sample. We then subjected aliquots to different processing methods (DNA extraction methods, internal standard addition, propidium monoazide (PMA) treatment, and whole-cell serial filtration). We evaluated the effects of these methods based on DNA yields and metagenomic sequencing outcomes. We also compared the metagenomic results to the microbial identifications obtained by cultivation using environmental microbiology methods and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS). Results: Our results demonstrate that bead-beating and heat lysis followed by liquid-liquid extraction is the optimal method for the identification of low-biomass surface-associated microbes, as opposed to widely used column-based and magnetic bead-based methods. For low-biomass surface-associated samples, ~590,000 reads per sample are sufficient for ≍90% coverage in metagenomic sequencing (Fig. 1). The ZymoBIOMICS microbial community standard is not appropriate for methods assessing membrane integrity. For the identification of putatively viable microorganisms, PMA treatment is promising, although elimination of signals from nonviable organisms will reduce the overall detectable signal. Combining PMA-treated metagenomic sequencing with cultivation yields the most comprehensive results, particularly for low-abundance taxa, despite high sequencing coverage (Fig. 2). To distribute more detection resources to bacteria, our target domain, we tried whole-cell filtration prior to extraction, attempting to isolate bacterial cells from eukaryotic cells and other particles. For low-biomass surface-associated samples, the sample loss and the difficulties in performing filtration outweigh the slight increase of bacterial signal. Conclusions: Despite optimization, we observed certain blind spots in both cultivation and metagenomic sequencing. This information is essential for informed risk assessment. Further research is needed to identify additional limitations to ensure that results from metagenomic sequencing can be interpreted in the context of healthcare-acquired infection prevention.
Funding: This work was supported by the Centers for Disease Control and Prevention (BAA FY2018-OADS-01 Contract 02915).
Disclosures: None
3343 Identification of host-microbial interaction networks that mediate intestinal epithelial barrier function in necrotizing enterocolitis
- David R Hill, Roberto Cieza, Veda K. Yadagiri, Phillip Tarr, Jason R. Spence, Vincent B. Young
-
- Journal:
- Journal of Clinical and Translational Science / Volume 3 / Issue s1 / March 2019
- Published online by Cambridge University Press:
- 26 March 2019, p. 13
-
- Article
-
- You have access Access
- Open access
- Export citation
-
OBJECTIVES/SPECIFIC AIMS: The central goal of this proposal is to characterize the mechanisms that mediate success or failure of immature intestinal barrier in necrotizing enterocilitis. METHODS/STUDY POPULATION: To do this, I will utilize stem cell derived human intestinal organoids (HIOs), an innovative model of the immature intestine, and a cohort of bacterial isolates collected from premature infants who developed NEC to interrogate the cause-effect relationship of these strains on maintenance of the intestinal barrier. I hypothesize that the epithelial response to bacterial colonization is strain-dependent and results in differences in inflammatory signaling that shape epithelial barrier function in the immature intestine. RESULTS/ANTICIPATED RESULTS: Preliminary data shows that colonization of HIOs with different bacteria leads to species-specific changes in barrier function, and some species selectively damage the epithelial barrier while others enhance epithelial barrier function. I have identified key inflammatory signals that serve as central drivers of intestinal barrier function. DISCUSSION/SIGNIFICANCE OF IMPACT: Characterization of this process is expected to substantially advance scientific understanding of early events in NEC pathogenesis and lead to new opportunities for targeted therapeutic intervention to accelerate barrier maturation or prevent hyperinflammatory reactivity in the neonatal intestine. The research proposed in this application represents an entirely novel approach to studying host-microbial interactions in the immature. Conceptually, this novel translational approach will help to define the pivotal role of colonizing bacteria in initiating epithelial inflammation in NEC patients.
3185 A Randomized Controlled Trial Comparing the Nonabsorbable Antibiotic Rifaximin vs. Dietary Intervention Low in Fermentable Sugars (FODMAP) in Irritable Bowel Syndrome
- Allen Lee, Krishna Rao, Emily Haller, Lauren Van Dam, Jason Baker, Shanti Eswaran, William Chey, Vincent Young, Chung Owyang, William Hasler
-
- Journal:
- Journal of Clinical and Translational Science / Volume 3 / Issue s1 / March 2019
- Published online by Cambridge University Press:
- 26 March 2019, p. 31
-
- Article
-
- You have access Access
- Open access
- Export citation
-
OBJECTIVES/SPECIFIC AIMS: Objectives and goals of this study are to (i) determine whether IBS-D patients randomized to either rifaximin or low FODMAP diet show improvement in IBS-related symptoms; and (2) identify using longitudinal analyses how SIBO status and fecal microbiota features associate with response to either rifaximin or low FODMAP dietary intervention. METHODS/STUDY POPULATION: 42 patients ≥ 18 years of age who meet Rome IV criteria for IBS-D will be randomized to receive either rifaximin or low FODMAP diet intervention. The primary outcome will be the proportion of responders to intervention which is defined as ≥ 30% reduction in mean daily abdominal pain or bloating by visual analog scale compared with baseline. Exclusion criteria will include: (a) history of microscopic colitis, inflammatory bowel disease, celiac disease, or other organic disease that could explain symptoms, (b) prior gastrointestinal surgery, other than appendectomy or cholecystectomy > 6 months prior to study initiation, (c) prior use of rifaximin or formal dietary interventions for IBS-D, (d) use of antibiotics within the past 3 months, or (e) use of probiotics within 1 month of study entry. Glucose hydrogen breath tests will be performed at the beginning and end of the trial to evaluate for SIBO. Fecal samples will be collected at 0, 2, and 6 weeks to determine changes in fecal microbial composition and structure. RESULTS/ANTICIPATED RESULTS: This study seeks to examine whether longitudinal analyses of small intestinal and colonic microbiota can subtype IBS-D subjects into clinically relevant phenotypes. A total of 18 subjects have been enrolled into the study. Clinical variables, hydrogen breath test results, and fecal microbiota data are being collected for ongoing analysis. DISCUSSION/SIGNIFICANCE OF IMPACT: Results from this study may help move treatment of IBS from a purely symptom based approach to a more individualized approach by stratifying IBS-D patients into distinct clinical phenotypes which are amenable to targeted therapeutic approaches.
A Generalizable, Data-Driven Approach to Predict Daily Risk of Clostridium difficile Infection at Two Large Academic Health Centers
- Jeeheh Oh, Maggie Makar, Christopher Fusco, Robert McCaffrey, Krishna Rao, Erin E. Ryan, Laraine Washer, Lauren R. West, Vincent B. Young, John Guttag, David C. Hooper, Erica S. Shenoy, Jenna Wiens
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 39 / Issue 4 / April 2018
- Published online by Cambridge University Press:
- 26 March 2018, pp. 425-433
- Print publication:
- April 2018
-
- Article
-
- You have access Access
- HTML
- Export citation
-
OBJECTIVE
An estimated 293,300 healthcare-associated cases of Clostridium difficile infection (CDI) occur annually in the United States. To date, research has focused on developing risk prediction models for CDI that work well across institutions. However, this one-size-fits-all approach ignores important hospital-specific factors. We focus on a generalizable method for building facility-specific models. We demonstrate the applicability of the approach using electronic health records (EHR) from the University of Michigan Hospitals (UM) and the Massachusetts General Hospital (MGH).
METHODSWe utilized EHR data from 191,014 adult admissions to UM and 65,718 adult admissions to MGH. We extracted patient demographics, admission details, patient history, and daily hospitalization details, resulting in 4,836 features from patients at UM and 1,837 from patients at MGH. We used L2 regularized logistic regression to learn the models, and we measured the discriminative performance of the models on held-out data from each hospital.
RESULTSUsing the UM and MGH test data, the models achieved area under the receiver operating characteristic curve (AUROC) values of 0.82 (95% confidence interval [CI], 0.80–0.84) and 0.75 ( 95% CI, 0.73–0.78), respectively. Some predictive factors were shared between the 2 models, but many of the top predictive factors differed between facilities.
CONCLUSIONA data-driven approach to building models for estimating daily patient risk for CDI was used to build institution-specific models at 2 large hospitals with different patient populations and EHR systems. In contrast to traditional approaches that focus on developing models that apply across hospitals, our generalizable approach yields risk-stratification models tailored to an institution. These hospital-specific models allow for earlier and more accurate identification of high-risk patients and better targeting of infection prevention strategies.
Infect Control Hosp Epidemiol 2018;39:425–433
Fecal Microbiota Therapy: Ready for Prime Time?
- Krishna Rao, Vincent B. Young, David M. Aronoff
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 35 / Issue 1 / January 2014
- Published online by Cambridge University Press:
- 10 May 2016, pp. 28-30
- Print publication:
- January 2014
-
- Article
- Export citation
Detection of Mixed Populations of Clostridium difficile from Symptomatic Patients Using Capillary-Based Polymerase Chain Reaction Ribotyping
- Adam A. Behroozian, Jeffrey P. Chludzinski, Eugene S. Lo, Sarah A. Ewing, Sheila Waslawski, Duane W. Newton, Vincent B. Young, David M. Aronoff, Seth T. Walk
-
- Journal:
- Infection Control & Hospital Epidemiology / Volume 34 / Issue 9 / September 2013
- Published online by Cambridge University Press:
- 02 January 2015, pp. 961-966
- Print publication:
- September 2013
-
- Article
- Export citation
-
Objective.
To investigate the simultaneous occurrence of more than 1 Clostridium difficile ribotype in patients' stool samples at the time of diagnostic testing.
Methods.Stool samples submitted for diagnostic testing for the presence of toxigenic C. difficile were obtained for 102 unique patients. A total of 95 single colonies of C. difficile per stool sample were isolated on selective media, subcultured alongside negative (uninoculated) controls, and polymerase chain reaction (PCR) ribotyped using capillary gel electrophoresis.
Results.Capillary-based PCR ribotyping was successful for 9,335 C. difficile isolates, yielding a median of 93 characterized isolates per stool sample (range, 69-95). More than 1 C. difficile ribotype was present in 16 of 102 (16%) C. difficile infection (CDI) cases; 2 of the 16 mixtures were composed of at least 3 ribotypes, while the remaining 14 were composed of at least 2.
Conclusions.Deep sampling of patient stool samples coupled with capillary-based PCR ribotyping identified a high rate of mixed CDI cases compared with previous estimates. Studies seeking to quantify the clinical significance of particular C. difficile ribotypes should account for mixed cases of disease.
Contributors
-
- By Rose Teteki Abbey, K. C. Abraham, David Tuesday Adamo, LeRoy H. Aden, Efrain Agosto, Victor Aguilan, Gillian T. W. Ahlgren, Charanjit Kaur AjitSingh, Dorothy B E A Akoto, Giuseppe Alberigo, Daniel E. Albrecht, Ruth Albrecht, Daniel O. Aleshire, Urs Altermatt, Anand Amaladass, Michael Amaladoss, James N. Amanze, Lesley G. Anderson, Thomas C. Anderson, Victor Anderson, Hope S. Antone, María Pilar Aquino, Paula Arai, Victorio Araya Guillén, S. Wesley Ariarajah, Ellen T. Armour, Brett Gregory Armstrong, Atsuhiro Asano, Naim Stifan Ateek, Mahmoud Ayoub, John Alembillah Azumah, Mercedes L. García Bachmann, Irena Backus, J. Wayne Baker, Mieke Bal, Lewis V. Baldwin, William Barbieri, António Barbosa da Silva, David Basinger, Bolaji Olukemi Bateye, Oswald Bayer, Daniel H. Bays, Rosalie Beck, Nancy Elizabeth Bedford, Guy-Thomas Bedouelle, Chorbishop Seely Beggiani, Wolfgang Behringer, Christopher M. Bellitto, Byard Bennett, Harold V. Bennett, Teresa Berger, Miguel A. Bernad, Henley Bernard, Alan E. Bernstein, Jon L. Berquist, Johannes Beutler, Ana María Bidegain, Matthew P. Binkewicz, Jennifer Bird, Joseph Blenkinsopp, Dmytro Bondarenko, Paulo Bonfatti, Riet en Pim Bons-Storm, Jessica A. Boon, Marcus J. Borg, Mark Bosco, Peter C. Bouteneff, François Bovon, William D. Bowman, Paul S. Boyer, David Brakke, Richard E. Brantley, Marcus Braybrooke, Ian Breward, Ênio José da Costa Brito, Jewel Spears Brooker, Johannes Brosseder, Nicholas Canfield Read Brown, Robert F. Brown, Pamela K. Brubaker, Walter Brueggemann, Bishop Colin O. Buchanan, Stanley M. Burgess, Amy Nelson Burnett, J. Patout Burns, David B. Burrell, David Buttrick, James P. Byrd, Lavinia Byrne, Gerado Caetano, Marcos Caldas, Alkiviadis Calivas, William J. Callahan, Salvatore Calomino, Euan K. Cameron, William S. Campbell, Marcelo Ayres Camurça, Daniel F. Caner, Paul E. Capetz, Carlos F. Cardoza-Orlandi, Patrick W. Carey, Barbara Carvill, Hal Cauthron, Subhadra Mitra Channa, Mark D. Chapman, James H. Charlesworth, Kenneth R. Chase, Chen Zemin, Luciano Chianeque, Philip Chia Phin Yin, Francisca H. Chimhanda, Daniel Chiquete, John T. Chirban, Soobin Choi, Robert Choquette, Mita Choudhury, Gerald Christianson, John Chryssavgis, Sejong Chun, Esther Chung-Kim, Charles M. A. Clark, Elizabeth A. Clark, Sathianathan Clarke, Fred Cloud, John B. Cobb, W. Owen Cole, John A Coleman, John J. Collins, Sylvia Collins-Mayo, Paul K. Conkin, Beth A. Conklin, Sean Connolly, Demetrios J. Constantelos, Michael A. Conway, Paula M. Cooey, Austin Cooper, Michael L. Cooper-White, Pamela Cooper-White, L. William Countryman, Sérgio Coutinho, Pamela Couture, Shannon Craigo-Snell, James L. Crenshaw, David Crowner, Humberto Horacio Cucchetti, Lawrence S. Cunningham, Elizabeth Mason Currier, Emmanuel Cutrone, Mary L. Daniel, David D. Daniels, Robert Darden, Rolf Darge, Isaiah Dau, Jeffry C. Davis, Jane Dawson, Valentin Dedji, John W. de Gruchy, Paul DeHart, Wendy J. Deichmann Edwards, Miguel A. De La Torre, George E. Demacopoulos, Thomas de Mayo, Leah DeVun, Beatriz de Vasconcellos Dias, Dennis C. Dickerson, John M. Dillon, Luis Miguel Donatello, Igor Dorfmann-Lazarev, Susanna Drake, Jonathan A. Draper, N. Dreher Martin, Otto Dreydoppel, Angelyn Dries, A. J. Droge, Francis X. D'Sa, Marilyn Dunn, Nicole Wilkinson Duran, Rifaat Ebied, Mark J. Edwards, William H. Edwards, Leonard H. Ehrlich, Nancy L. Eiesland, Martin Elbel, J. Harold Ellens, Stephen Ellingson, Marvin M. Ellison, Robert Ellsberg, Jean Bethke Elshtain, Eldon Jay Epp, Peter C. Erb, Tassilo Erhardt, Maria Erling, Noel Leo Erskine, Gillian R. Evans, Virginia Fabella, Michael A. Fahey, Edward Farley, Margaret A. Farley, Wendy Farley, Robert Fastiggi, Seena Fazel, Duncan S. Ferguson, Helwar Figueroa, Paul Corby Finney, Kyriaki Karidoyanes FitzGerald, Thomas E. FitzGerald, John R. Fitzmier, Marie Therese Flanagan, Sabina Flanagan, Claude Flipo, Ronald B. Flowers, Carole Fontaine, David Ford, Mary Ford, Stephanie A. Ford, Jim Forest, William Franke, Robert M. Franklin, Ruth Franzén, Edward H. Friedman, Samuel Frouisou, Lorelei F. Fuchs, Jojo M. Fung, Inger Furseth, Richard R. Gaillardetz, Brandon Gallaher, China Galland, Mark Galli, Ismael García, Tharscisse Gatwa, Jean-Marie Gaudeul, Luis María Gavilanes del Castillo, Pavel L. Gavrilyuk, Volney P. Gay, Metropolitan Athanasios Geevargis, Kondothra M. George, Mary Gerhart, Simon Gikandi, Maurice Gilbert, Michael J. Gillgannon, Verónica Giménez Beliveau, Terryl Givens, Beth Glazier-McDonald, Philip Gleason, Menghun Goh, Brian Golding, Bishop Hilario M. Gomez, Michelle A. Gonzalez, Donald K. Gorrell, Roy Gottfried, Tamara Grdzelidze, Joel B. Green, Niels Henrik Gregersen, Cristina Grenholm, Herbert Griffiths, Eric W. Gritsch, Erich S. Gruen, Christoffer H. Grundmann, Paul H. Gundani, Jon P. Gunnemann, Petre Guran, Vidar L. Haanes, Jeremiah M. Hackett, Getatchew Haile, Douglas John Hall, Nicholas Hammond, Daphne Hampson, Jehu J. Hanciles, Barry Hankins, Jennifer Haraguchi, Stanley S. Harakas, Anthony John Harding, Conrad L. Harkins, J. William Harmless, Marjory Harper, Amir Harrak, Joel F. Harrington, Mark W. Harris, Susan Ashbrook Harvey, Van A. Harvey, R. Chris Hassel, Jione Havea, Daniel Hawk, Diana L. Hayes, Leslie Hayes, Priscilla Hayner, S. Mark Heim, Simo Heininen, Richard P. Heitzenrater, Eila Helander, David Hempton, Scott H. Hendrix, Jan-Olav Henriksen, Gina Hens-Piazza, Carter Heyward, Nicholas J. Higham, David Hilliard, Norman A. Hjelm, Peter C. Hodgson, Arthur Holder, M. Jan Holton, Dwight N. Hopkins, Ronnie Po-chia Hsia, Po-Ho Huang, James Hudnut-Beumler, Jennifer S. Hughes, Leonard M. Hummel, Mary E. Hunt, Laennec Hurbon, Mark Hutchinson, Susan E. Hylen, Mary Beth Ingham, H. Larry Ingle, Dale T. Irvin, Jon Isaak, Paul John Isaak, Ada María Isasi-Díaz, Hans Raun Iversen, Margaret C. Jacob, Arthur James, Maria Jansdotter-Samuelsson, David Jasper, Werner G. Jeanrond, Renée Jeffery, David Lyle Jeffrey, Theodore W. Jennings, David H. Jensen, Robin Margaret Jensen, David Jobling, Dale A. Johnson, Elizabeth A. Johnson, Maxwell E. Johnson, Sarah Johnson, Mark D. Johnston, F. Stanley Jones, James William Jones, John R. Jones, Alissa Jones Nelson, Inge Jonsson, Jan Joosten, Elizabeth Judd, Mulambya Peggy Kabonde, Robert Kaggwa, Sylvester Kahakwa, Isaac Kalimi, Ogbu U. Kalu, Eunice Kamaara, Wayne C. Kannaday, Musimbi Kanyoro, Veli-Matti Kärkkäinen, Frank Kaufmann, Léon Nguapitshi Kayongo, Richard Kearney, Alice A. Keefe, Ralph Keen, Catherine Keller, Anthony J. Kelly, Karen Kennelly, Kathi Lynn Kern, Fergus Kerr, Edward Kessler, George Kilcourse, Heup Young Kim, Kim Sung-Hae, Kim Yong-Bock, Kim Yung Suk, Richard King, Thomas M. King, Robert M. Kingdon, Ross Kinsler, Hans G. Kippenberg, Cheryl A. Kirk-Duggan, Clifton Kirkpatrick, Leonid Kishkovsky, Nadieszda Kizenko, Jeffrey Klaiber, Hans-Josef Klauck, Sidney Knight, Samuel Kobia, Robert Kolb, Karla Ann Koll, Heikki Kotila, Donald Kraybill, Philip D. W. Krey, Yves Krumenacker, Jeffrey Kah-Jin Kuan, Simanga R. Kumalo, Peter Kuzmic, Simon Shui-Man Kwan, Kwok Pui-lan, André LaCocque, Stephen E. Lahey, John Tsz Pang Lai, Emiel Lamberts, Armando Lampe, Craig Lampe, Beverly J. Lanzetta, Eve LaPlante, Lizette Larson-Miller, Ariel Bybee Laughton, Leonard Lawlor, Bentley Layton, Robin A. Leaver, Karen Lebacqz, Archie Chi Chung Lee, Marilyn J. Legge, Hervé LeGrand, D. L. LeMahieu, Raymond Lemieux, Bill J. Leonard, Ellen M. Leonard, Outi Leppä, Jean Lesaulnier, Nantawan Boonprasat Lewis, Henrietta Leyser, Alexei Lidov, Bernard Lightman, Paul Chang-Ha Lim, Carter Lindberg, Mark R. Lindsay, James R. Linville, James C. Livingston, Ann Loades, David Loades, Jean-Claude Loba-Mkole, Lo Lung Kwong, Wati Longchar, Eleazar López, David W. Lotz, Andrew Louth, Robin W. Lovin, William Luis, Frank D. Macchia, Diarmaid N. J. MacCulloch, Kirk R. MacGregor, Marjory A. MacLean, Donald MacLeod, Tomas S. Maddela, Inge Mager, Laurenti Magesa, David G. Maillu, Fortunato Mallimaci, Philip Mamalakis, Kä Mana, Ukachukwu Chris Manus, Herbert Robinson Marbury, Reuel Norman Marigza, Jacqueline Mariña, Antti Marjanen, Luiz C. L. Marques, Madipoane Masenya (ngwan'a Mphahlele), Caleb J. D. Maskell, Steve Mason, Thomas Massaro, Fernando Matamoros Ponce, András Máté-Tóth, Odair Pedroso Mateus, Dinis Matsolo, Fumitaka Matsuoka, John D'Arcy May, Yelena Mazour-Matusevich, Theodore Mbazumutima, John S. McClure, Christian McConnell, Lee Martin McDonald, Gary B. McGee, Thomas McGowan, Alister E. McGrath, Richard J. McGregor, John A. McGuckin, Maud Burnett McInerney, Elsie Anne McKee, Mary B. McKinley, James F. McMillan, Ernan McMullin, Kathleen E. McVey, M. Douglas Meeks, Monica Jyotsna Melanchthon, Ilie Melniciuc-Puica, Everett Mendoza, Raymond A. Mentzer, William W. Menzies, Ina Merdjanova, Franziska Metzger, Constant J. Mews, Marvin Meyer, Carol Meyers, Vasile Mihoc, Gunner Bjerg Mikkelsen, Maria Inêz de Castro Millen, Clyde Lee Miller, Bonnie J. Miller-McLemore, Alexander Mirkovic, Paul Misner, Nozomu Miyahira, R. W. L. Moberly, Gerald Moede, Aloo Osotsi Mojola, Sunanda Mongia, Rebeca Montemayor, James Moore, Roger E. Moore, Craig E. Morrison O.Carm, Jeffry H. Morrison, Keith Morrison, Wilson J. Moses, Tefetso Henry Mothibe, Mokgethi Motlhabi, Fulata Moyo, Henry Mugabe, Jesse Ndwiga Kanyua Mugambi, Peggy Mulambya-Kabonde, Robert Bruce Mullin, Pamela Mullins Reaves, Saskia Murk Jansen, Heleen L. Murre-Van den Berg, Augustine Musopole, Isaac M. T. Mwase, Philomena Mwaura, Cecilia Nahnfeldt, Anne Nasimiyu Wasike, Carmiña Navia Velasco, Thulani Ndlazi, Alexander Negrov, James B. Nelson, David G. Newcombe, Carol Newsom, Helen J. Nicholson, George W. E. Nickelsburg, Tatyana Nikolskaya, Damayanthi M. A. Niles, Bertil Nilsson, Nyambura Njoroge, Fidelis Nkomazana, Mary Beth Norton, Christian Nottmeier, Sonene Nyawo, Anthère Nzabatsinda, Edward T. Oakes, Gerald O'Collins, Daniel O'Connell, David W. Odell-Scott, Mercy Amba Oduyoye, Kathleen O'Grady, Oyeronke Olajubu, Thomas O'Loughlin, Dennis T. Olson, J. Steven O'Malley, Cephas N. Omenyo, Muriel Orevillo-Montenegro, César Augusto Ornellas Ramos, Agbonkhianmeghe E. Orobator, Kenan B. Osborne, Carolyn Osiek, Javier Otaola Montagne, Douglas F. Ottati, Anna May Say Pa, Irina Paert, Jerry G. Pankhurst, Aristotle Papanikolaou, Samuele F. Pardini, Stefano Parenti, Peter Paris, Sung Bae Park, Cristián G. Parker, Raquel Pastor, Joseph Pathrapankal, Daniel Patte, W. Brown Patterson, Clive Pearson, Keith F. Pecklers, Nancy Cardoso Pereira, David Horace Perkins, Pheme Perkins, Edward N. Peters, Rebecca Todd Peters, Bishop Yeznik Petrossian, Raymond Pfister, Peter C. Phan, Isabel Apawo Phiri, William S. F. Pickering, Derrick G. Pitard, William Elvis Plata, Zlatko Plese, John Plummer, James Newton Poling, Ronald Popivchak, Andrew Porter, Ute Possekel, James M. Powell, Enos Das Pradhan, Devadasan Premnath, Jaime Adrían Prieto Valladares, Anne Primavesi, Randall Prior, María Alicia Puente Lutteroth, Eduardo Guzmão Quadros, Albert Rabil, Laurent William Ramambason, Apolonio M. Ranche, Vololona Randriamanantena Andriamitandrina, Lawrence R. Rast, Paul L. Redditt, Adele Reinhartz, Rolf Rendtorff, Pål Repstad, James N. Rhodes, John K. Riches, Joerg Rieger, Sharon H. Ringe, Sandra Rios, Tyler Roberts, David M. Robinson, James M. Robinson, Joanne Maguire Robinson, Richard A. H. Robinson, Roy R. Robson, Jack B. Rogers, Maria Roginska, Sidney Rooy, Rev. Garnett Roper, Maria José Fontelas Rosado-Nunes, Andrew C. Ross, Stefan Rossbach, François Rossier, John D. Roth, John K. Roth, Phillip Rothwell, Richard E. Rubenstein, Rosemary Radford Ruether, Markku Ruotsila, John E. Rybolt, Risto Saarinen, John Saillant, Juan Sanchez, Wagner Lopes Sanchez, Hugo N. Santos, Gerhard Sauter, Gloria L. Schaab, Sandra M. Schneiders, Quentin J. Schultze, Fernando F. Segovia, Turid Karlsen Seim, Carsten Selch Jensen, Alan P. F. Sell, Frank C. Senn, Kent Davis Sensenig, Damían Setton, Bal Krishna Sharma, Carolyn J. Sharp, Thomas Sheehan, N. Gerald Shenk, Christian Sheppard, Charles Sherlock, Tabona Shoko, Walter B. Shurden, Marguerite Shuster, B. Mark Sietsema, Batara Sihombing, Neil Silberman, Clodomiro Siller, Samuel Silva-Gotay, Heikki Silvet, John K. Simmons, Hagith Sivan, James C. Skedros, Abraham Smith, Ashley A. Smith, Ted A. Smith, Daud Soesilo, Pia Søltoft, Choan-Seng (C. S.) Song, Kathryn Spink, Bryan Spinks, Eric O. Springsted, Nicolas Standaert, Brian Stanley, Glen H. Stassen, Karel Steenbrink, Stephen J. Stein, Andrea Sterk, Gregory E. Sterling, Columba Stewart, Jacques Stewart, Robert B. Stewart, Cynthia Stokes Brown, Ken Stone, Anne Stott, Elizabeth Stuart, Monya Stubbs, Marjorie Hewitt Suchocki, David Kwang-sun Suh, Scott W. Sunquist, Keith Suter, Douglas Sweeney, Charles H. Talbert, Shawqi N. Talia, Elsa Tamez, Joseph B. Tamney, Jonathan Y. Tan, Yak-Hwee Tan, Kathryn Tanner, Feiya Tao, Elizabeth S. Tapia, Aquiline Tarimo, Claire Taylor, Mark Lewis Taylor, Bishop Abba Samuel Wolde Tekestebirhan, Eugene TeSelle, M. Thomas Thangaraj, David R. Thomas, Andrew Thornley, Scott Thumma, Marcelo Timotheo da Costa, George E. “Tink” Tinker, Ola Tjørhom, Karen Jo Torjesen, Iain R. Torrance, Fernando Torres-Londoño, Archbishop Demetrios [Trakatellis], Marit Trelstad, Christine Trevett, Phyllis Trible, Johannes Tromp, Paul Turner, Robert G. Tuttle, Archbishop Desmond Tutu, Peter Tyler, Anders Tyrberg, Justin Ukpong, Javier Ulloa, Camillus Umoh, Kristi Upson-Saia, Martina Urban, Monica Uribe, Elochukwu Eugene Uzukwu, Richard Vaggione, Gabriel Vahanian, Paul Valliere, T. J. Van Bavel, Steven Vanderputten, Peter Van der Veer, Huub Van de Sandt, Louis Van Tongeren, Luke A. Veronis, Noel Villalba, Ramón Vinke, Tim Vivian, David Voas, Elena Volkova, Katharina von Kellenbach, Elina Vuola, Timothy Wadkins, Elaine M. Wainwright, Randi Jones Walker, Dewey D. Wallace, Jerry Walls, Michael J. Walsh, Philip Walters, Janet Walton, Jonathan L. Walton, Wang Xiaochao, Patricia A. Ward, David Harrington Watt, Herold D. Weiss, Laurence L. Welborn, Sharon D. Welch, Timothy Wengert, Traci C. West, Merold Westphal, David Wetherell, Barbara Wheeler, Carolinne White, Jean-Paul Wiest, Frans Wijsen, Terry L. Wilder, Felix Wilfred, Rebecca Wilkin, Daniel H. Williams, D. Newell Williams, Michael A. Williams, Vincent L. Wimbush, Gabriele Winkler, Anders Winroth, Lauri Emílio Wirth, James A. Wiseman, Ebba Witt-Brattström, Teofil Wojciechowski, John Wolffe, Kenman L. Wong, Wong Wai Ching, Linda Woodhead, Wendy M. Wright, Rose Wu, Keith E. Yandell, Gale A. Yee, Viktor Yelensky, Yeo Khiok-Khng, Gustav K. K. Yeung, Angela Yiu, Amos Yong, Yong Ting Jin, You Bin, Youhanna Nessim Youssef, Eliana Yunes, Robert Michael Zaller, Valarie H. Ziegler, Barbara Brown Zikmund, Joyce Ann Zimmerman, Aurora Zlotnik, Zhuo Xinping
- Edited by Daniel Patte, Vanderbilt University, Tennessee
-
- Book:
- The Cambridge Dictionary of Christianity
- Published online:
- 05 August 2012
- Print publication:
- 20 September 2010, pp xi-xliv
-
- Chapter
- Export citation
Contributors
-
- By Joanne R. Adler, David A. Alexander, Laurence Alison, Catherine C. Ayoub, Peter Banister, Anthony R. Beech, Amanda Biggs, Julian Boon, Adrian Bowers, Neil Brewer, Eric Broekaert, Paula Brough, Jennifer M. Brown, Kevin Browne, Elizabeth A. Campbell, David Canter, Michael Carlin, Shihning Chou, Martin A. Conway, Claire Cooke, David Cooke, Ilse Derluyn, Robert J. Edelmann, Vincent Egan, Tom Ellis, Marie Eyre, David P. Farrington, Seena Fazel, Daniel B. Fishman, Victoria Follette, Katarina Fritzon, Elizabeth Gilchrist, Nathan D. Gillard, Renée Gobeil, Agnieszka Golec de Zavala, Jane Goodman-Delahunty, Lynsey Gozna, Don Grubin, Gisli H. Gudjonsson, Helinä Häkkänen-Nyholm, Guy Hall, Nathan Hall, Roisin Hall, Sean Hammond, Leigh Harkins, Grant T. Harris, Camilla Herbert, Robert D. Hoge, Todd E. Hogue, Clive R. Hollin, Lorraine Hope, Miranda A. H. Horvath, Kevin Howells, Carol A. Ireland, Jane L. Ireland, Mark Kebbell, Michael King, Bruce D. Kirkcaldy, Heidi La Bash, Cara Laney, William R. Lindsay, Elizabeth F. Loftus, L. E. Marshall, W. L. Marshall, James McGuire, Neil McKeganey, T. M. McMillan, Mary McMurran, Joav Merrick, Becky Milne, Joanne M. Nadkarni, Claire Nee, M. D. O’Brien, William O’Donohue, Darragh O’Neill, Jane Palmer, Adria Pearson, Derek Perkins, Devon L. L. Polaschek, Louise E. Porter, Charlotte C. Powell, Graham E. Powell, Martine Powell, Christine Puckering, Ethel Quayle, Vernon L. Quinsey, Marnie E. Rice, Randall Richardson-Vejlgaard, Richard Rogers, Louis B Schlesinger, Carolyn Semmler, G. A. Serran, Ralph C. Serin, John L. Taylor, Max Taylor, Brian Thomas-Peter, Paul A. Tiffin, Graham Towl, Rosie Travers, Arlene Vetere, Graham Wagstaff, Helen Wakeling, Fiona Warren, Brandon C. Welsh, David Wexler, Margaret Wilson, Dan Yarmey, Susan Young
- Edited by Jennifer M. Brown, London School of Economics and Political Science, Elizabeth A. Campbell, University of Glasgow
-
- Book:
- The Cambridge Handbook of Forensic Psychology
- Published online:
- 06 July 2010
- Print publication:
- 29 April 2010, pp xix-xxiii
-
- Chapter
- Export citation
Explaining unexplained diarrhea and associating risks and infections
- Donna M. Denno, Eileen J. Klein, Vincent B. Young, James G. Fox, David Wang, Phillip I. Tarr
-
- Journal:
- Animal Health Research Reviews / Volume 8 / Issue 1 / June 2007
- Published online by Cambridge University Press:
- 13 August 2007, pp. 69-80
-
- Article
- Export citation
-
Gastrointestinal illnesses are common afflictions. However, knowledge of their etiology is often lacking. Moreover, most cases of infections with reportable enteric pathogens (Campylobacter jejuni, Escherichia coli O157:H7, Salmonella, Shigella, Yersinia, Cryptosporidia and Giardia) have sporadic modes of acquisition, yet control measures are often biased towards mitigation of risks discerned by outbreak analysis. To determine the etiology of unexplained diarrhea it is important to study populations that can be matched to appropriate controls and to couple thorough classic microbiologic evaluation on receipt of specimens with archiving and outgrowth capabilities. Research evaluations should address the potential roles of a broad panel of candidate bacterial pathogens including diarrheagenic E. coli, Listeria monocytogenes, Helicobacters and jejuni Campylobacters, and also apply novel massively parallel sequencing and nucleic acid detection technologies that allow the detection of viral pathogens. To fill voids in our knowledge regarding sources of known enteric pathogens it will be critical to extend case-control studies to assess risk factors and exposures to patients with non-epidemic illnesses and to appropriate controls. By filling these gaps in our knowledge it should be possible to formulate rational prevention mechanisms for human gastrointestinal illnesses.




